With the psfmi_mm function pooling and selection of (generalized) linear mixed models, i.e linear and logistic mixed models, is possible. With the psfmi_stab function the stability of models after using psfmi_lr, psfmi_coxr and psfmi_mm can be evaluated. The latter function uses (single) bootstrapping for the psfmi_lr and psfmi_coxr functions and cluster bootstrapping for the psfmi_mm function. With the function the bootstrap inclusion frequency of predictors and models can be estimated.

More information about the package can be found on the Cran website.
You can easily install the package by running install.packages("psfmi") in the Console window in Rstudio or R. The development version can be installed from Github by using: install.packages("devtools")
library(devtools)
devtools::install_github("mwheymans/psfmi")
library(psfmi)
If you have questions about the psfmi package send an email to mw.heymans@amsterdamumc.nl or leave a message below. Enjoy using the package!

I have written an online Applied Missing Data book for all researchers that have missing data. Click on continue reading to access the book.

With the psfmi package it is possible to force variables in the model during backward selection.
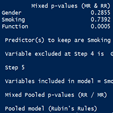
With the psfmi package it is possible to do backward selection for logistic regression models after Multiple Imputation (MI). For backward selection, several variable selection criteria can be used. These criteria are called the D1, D2, D3 and Median P-rule (MPR).

This post is about different functions of the survival package, like basehaz, predict.coxph, survfit, termplot, what kind of results they provide and how they are calculated.

